Hello, everyone!
We’d like to periodically share our progress over this analysis as there are lots of possible ways to advance over what’s available, it’s very useful to share ideas, get some feedback and also know, compare and test each others hypotheses.
Repository: https://gitlab.com/our-sci/rfc-docs/-/tree/master
Results (download and open in web browser): https://gitlab.com/our-sci/rfc-docs/-/raw/master/2019/dataAnalysis.html?inline=false
Our results would be mostly available in the dataAnalysis.html file inside our gitlab repo, where you can find associated R objects also.
We have begun with some fundamental steps to make the data available, mostly merging, cleaning, formatting and elimination of outliers. The data is now organized as can be seen in our spectral curves plots. The curves have been clustered by derivative sign successions in order to spot the most obvious patterns. In these plots we observed some curious behaviour which we are still analyzing but for now we mostly classified the most peculiar patterns as atypical data.
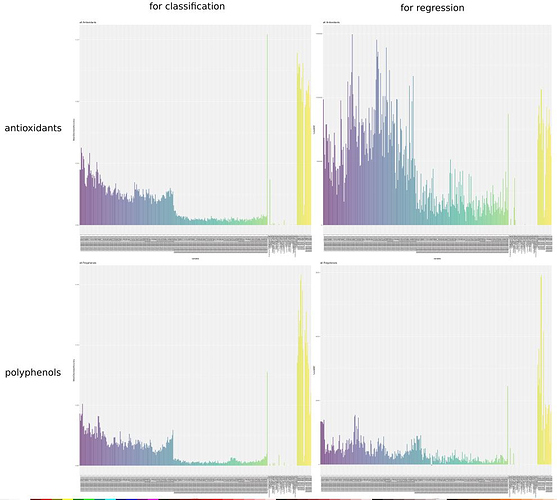
Our first target is to predict Polyphenols and Antioxidants contents using data obtained through the Our Sci’s spectrometer. We’ve already done exhaustive modelling work, over a great number of subsets of available dimensions (mostly combinations of OurSci’s spectra, the SWIR spectra measured with the SIWARE device and metadata as produce color) with a variety of available statistical models. We’ve organized a powerful testing framework which enabled us to run hundreds of thousands of modelling instances, with repetitions aimed at reasonable levels of cross validations ( 15 folds ) in order to avoid bias and overfit as much as possible. If you’d like us to run some particular model you believe would be suited for this situation and we haven’t tried, please let us know, we probably can run it using the same procedure without a great effort.
This is still a testing stage in which we are searching for the most promising models. This is the reason you’ll find some unreasonable models inside the tables, for example models that are clearly overfitted. This also allowed us to see, for example, that our cross validation was effectively differentiating overfitted linear models with ease (you can see this looking at the original rsquared column in the linear models table, in comparison to the cvRsquared one).
Special effort has been made to fit the classical linear models, first shown in the brief in a tentative ANCOVA fit, then detailed over the modelling section. A variance stabilizing transformation has been tried (without interesting outcomes) and a shallow essay over robust modelling has been carried, this one with an interesting reduction of 20% over the variance, without any special tuning. Robust fitting and outliers weighting is one of the directions we will explore further.
We’ve done preliminary work over deep learning models such as Neural Networks and Support Vector Machines, calibrating the same system we used for massive modelling iteration and looking for reasonable tuning ranges for the hyperparameters. We’ll offer a similar broad survey over these models. As involved calculations are exceedingly heavy, with these sections we started a new organization where the heavy modelling work will be done on separate files. This one is not finished and is still unmerged with the main document, in a separate modelling folder on the same repo.
We originally anounced promising results on our Random Forest models, this was mostly due to a failure in distinguishing repeated measurings in the dataset when doing cross validation. Event though they still show a superior fit than linear models, neither of them offers a working solution.
Up to this moment, no modelling solution has shown a conclusive, practically useful result . We’ve gone back gone back to check at the state of the art in similar problems and decided Functional Data Analysis seems to be the most promising solution. The expertise gained in working with this particular dataset and the iterative tidying and organizing steps these investigations have required have left us well equipped to test for further hypotheses and proceed with this new modelling options in shorter time, so hopefully we’ll soon see if this path is more fruitful.
These are some of the steps we’ll take during this week, until this Friday
- Finish a new round of outlier detection, searching also for “shape” outliers, that is curves that are between the acceptable ranges but have an unusual geometry/behaviuor. In general, we will attempt a better curing of the curves sets, we’ll try to explain the bottom clustering observed on most datasets and asses if it’s reasonable to attempt normalization (profiting on the existence of shared heavy absorption bands) or if we should split those sets over different models.
- Try functional regression solutions and assess their precision through a consistent validation framework. Recent literature suggests that this is the better suited theoretical framework.
- We’ll resort more systematically to available metadata, especially Brix content and produce color. We’ll see if dry mass percentage adds considerable predictive power, taking into account it’s less practical to implement for massive use of the model. We are also checking if processing time explains some of the unsual patterns.
- We’ll try to increase the power of our quantile predictions by increasing the number of quantiles and splitting into three bands (inferior, center and superior one).
We are looking forward to also hearing from your findings and observations and wish you luck in your own investigations.
 . Basically - we just need to hand those numbers to potential partners, bfa members and others and say ‘well… this is what we can do… is this useful?’.
. Basically - we just need to hand those numbers to potential partners, bfa members and others and say ‘well… this is what we can do… is this useful?’.